Modeling
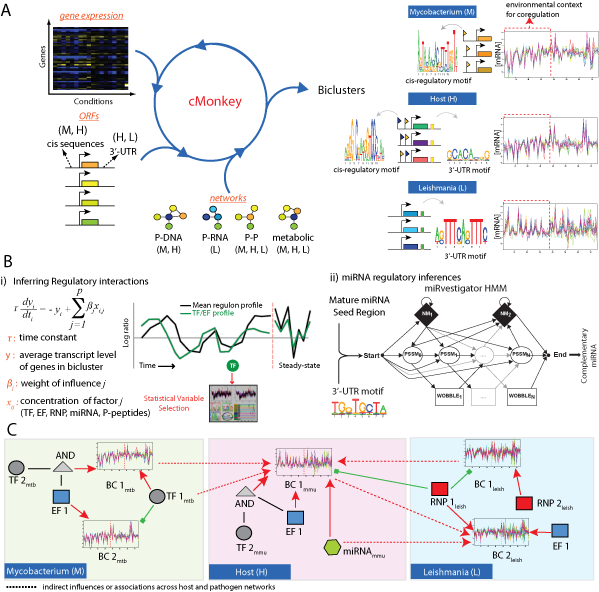
Overview of modeling algorithm components and proposed integrated host/pathogen model. cMonkey. (A) identifies modules of co-regulated genes under subsets of experimental conditions, by integration of gene expression data, de novo detection of cis-regulatory or 3’UTR motifs, and functional or physical association networks between component genes. Inferelator and miRvestigator (B) are complementary methods for inferring regulatory factor (TF and EF) or miRNA influences, respectively, on the modules produced by cMonkey. Inferelator uses constrained linear regression on an approximation to the time-dynamic expression for gene regulation to infer the most likely activating or repressing TFs or EFs for each module. miRvestigator applies a HMM to the de novo 3’UTR motifs detected during biclustering by cMonkey to estimate which, if any, known miRNA seed sequence is a strong match, and therefore a putative miRNA regulator. These methods will be integrated across host and pathogen to infer both individual host/pathogen networks, as well as putative influences between the species (C). See the Methods section for more details.
Tools





